Computational and Bioinformatics Approaches for Identifying Comorbidities of COVID-19 Using Transcriptomic Data
 , Md Mohiuddin2
, Md Mohiuddin2 , Salma Akhter3
, Salma Akhter3 , Md. Hasan Imam1
, Md. Hasan Imam1 , A. K. M. Mostofa Kamal Habib4
, A. K. M. Mostofa Kamal Habib4
Journal of Engineering Research and Sciences, Volume 3, Issue 4, Page # 32-41, 2024; DOI: 10.55708/js0304004
Keywords: COVID-19, Comorbidity Identification, Transcriptomic Data, Tippett’s Method, Euclidean Distance
(This article belongs to the Section Mathematical and Computational Biology (MCB))
Export Citations
Cite
Omit, H. B. S. , Mohiuddin, M. , Akhter, S. , Imam, M. H. , Habib, A. K. M. M. K. , Hossain, S. M. M. and Podder, N. K. (2024). Computational and Bioinformatics Approaches for Identifying Comorbidities of COVID-19 Using Transcriptomic Data. Journal of Engineering Research and Sciences, 3(4), 32–41. https://doi.org/10.55708/js0304004
hudeb Babu Sen Omit, Md Mohiuddin, Salma Akhter, Md. Hasan Imam, A. K. M. Mostofa Kamal Habib, Syed Mohammad Meraz Hossain and Nitun Kumar Podder. "Computational and Bioinformatics Approaches for Identifying Comorbidities of COVID-19 Using Transcriptomic Data." Journal of Engineering Research and Sciences 3, no. 4 (April 2024): 32–41. https://doi.org/10.55708/js0304004
H.B.S. Omit, M. Mohiuddin, S. Akhter, M.H. Imam, A.K.M.M.K. Habib, S.M.M. Hossain and N.K. Podder, "Computational and Bioinformatics Approaches for Identifying Comorbidities of COVID-19 Using Transcriptomic Data," Journal of Engineering Research and Sciences, vol. 3, no. 4, pp. 32–41, Apr. 2024, doi: 10.55708/js0304004.
Comorbidity is the co-existence of one or more diseases that occur concurrently or after the primary disease. Patients may have developed comorbidities for COVID-19 that cause harm to the patient’s organs. Besides, patients with existing comorbidities are at high risk, since mortality rates are strongly influenced by comorbidities or former health conditions. Therefore, we developed a computational and bioinformatics model to identify the comorbidities of COVID-19 utilizing transcriptome datasets of patient’s whole blood cells. In our model, we employed gene expression analysis to identify dysregulated genes and curated diseases from Gold Benchmark databases using the dysregulated genes. Subsequently, Tippett's Method is used for COVID-19’s P-value calculation, and according to the P-value, Euclidean distances are calculated between COVID-19 and the collected diseases. Then the collected diseases are ordered and clustered based on the Euclidean distance. Finally, comorbidities are selected from the top clusters based on a comprehensive literature search. Applying the model, we found that acute myelocytic leukemia, cancer of urinary tract, body weight changes, abdominal aortic aneurysm, kidney neoplasm, diabetes mellitus, and some other rare diseases have correlation with COVID-19 and many of them reveal as comorbidity. Since comorbidities are in conjunction with the primary disease, thus similar drugs and treatments can be used to recover both COVID-19 and its comorbidities by further research. We also proposed that this model can be further useful for detecting comorbidities of other diseases as well.
1. Introduction
COVID-19 is an infectious disease first reported in December 2019 in Wuhan, Hubei Province, China, caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Through direct contact or droplets from the infected person’s respiratory system, COVID-19 is spread between individuals [1], [2]. Consequently, COVID-19 has spread to almost all countries around the world, and on 11 March 2020, the World Health Organization (WHO) declared it a global pandemic. The WHO estimates that COVID-19 causes more than 180 million confirmed cases and 4 million deaths worldwide. This death is often caused by the underlying comorbidities of COVID-19, and the severity of the disease is increased by these existing conditions, as the comorbidities are correlated with morbidity and mortality [3] — [5]. In addition, COVID-19 indirectly impacts damage to organ systems and organs of the human body, which is also resulting in death [6] — [9]. It is therefore vital to develop a computation-based approach that can detect the comorbidities of COVID-19 so that early protection, prevention, and treatment can be implemented.
There have been several studies performed to identify COVID-19 comorbidities. In [10], the authors found that hypertension, diabetes, cardiovascular diseases, and chronic kidney disease are the most common comorbidities of COVID-19. In [11], cardiovascular disease and chronic respiratory disease are the riskiest comorbidities observed by the authors. Another study performed in [12] and the authors showed that hypertension, diabetes, obstructive pulmonary disease, cardiovascular disease, and cerebrovascular disease are the risk factors associated with COVID-19. Moreover, according to [13], hypertension, chronic cardiovascular disease, diabetes, and chronic cerebrovascular disease have a high percentage in COVID-19 patients. Besides, in [14], the authors reported hypertension, diabetes mellitus, cerebrovascular, cardiovascular disease, kidney disease, etc. as comorbidities of COVID-19. A similar analysis by the authors of [15] also reported that hypertension, diabetes, obesity, asthma, etc. are the comorbidities in hospitalized COVID-19 patients.
We found that a considerable number of works identifying the comorbidities of COVID-19 are based on literature-driven meta-analyses and some of them are discussed in the literature review section. Therefore, we have performed a computational and bioinformatics-based approach to detect COVID-19’s comorbidities
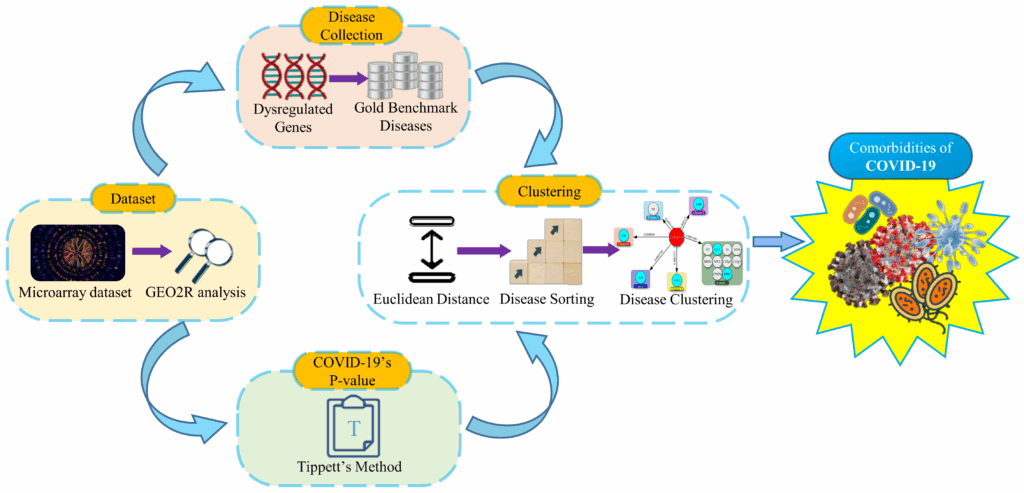
Our analysis was based on transcriptomic data derived from blood samples of COVID-19 patients and healthy individuals. Then we analyzed the dataset with the GEO2R (https://www.ncbi.nlm.nih.gov/geo/geo2r/) tool and identified up and down-regulated genes. As a result of these dysregulated genes, we compiled a list of associated diseases from Gold Benchmark databases. Then the P-value of COVID-19 is calculated, as described in the following sections. After that, Euclidean distance is used to measure how close the collected diseases are to COVID-19. The Euclidean distances are calculated between COVID-19’s P-value and the Gold Benchmark diseases’ P-value. Consequently, the short Euclidean distances mean more proximity of diseases with COVID-19. Next, we organized the collected diseases in small to large orders according to the Euclidean distance. Then the diseases of similar Euclidean distance are clustered from top to bottom of the organized list. In our final step, we selected the comorbidity from the top clusters.
The main contributions of our paper are as follows:
- As a result of conducting our analysis on genetic data, we avoided the influences of literature-driven type biases.
- As we have used a transcriptomic dataset of the whole genome from the blood sample, some novel findings can potentially be revealed as the comorbidity of COVID-19.
- Our pipeline is capable of generating automated results provided input dataset.
- Finally, if the transcriptomic dataset exists, it is possible to apply the technique swiftly to different diseases.
The rest of the paper is structured as follows: Section II describes related previous works and their shortcomings; Section III depicts the proposed pipeline outlining our methodology and analysis. Section IV discusses the outcomes and validation. Section V concludes our paper and introduces future extensions of the work.
2. Literature Review
According to the World Health Organization, there have been 170,426,245 confirmed cases, including 3,548,628 deaths due to the COVID-19 pandemic globally as of 1 June 2021. The number of infected people and deaths is increasing rapidly and this mortality, infection, and severity are getting more dangerous because of existing comorbidities and underlying health conditions [16] — [18]. Hence it is important to identify the comorbidities or risk factors to reduce severe COVID-19 progression. Some COVID-19 comorbidities associated research works are presented as follows:
A meta-analysis of the published global literature was conducted by the authors of [19] by combining clinical data with comorbidity details, they were able to identify significant COVID-19 comorbidities. They found chronic kidney disease, type-2 diabetes, malignancy, hypertension, cardiovascular disease, etc. as the most significant comorbidities associated with COVID-19; working with published articles and aggregated clinical cohort data there is a possibility of biases.
By reviewing the literature the authors of [20] selected angiotensin-converting enzyme 2 (ACE2), transmembrane protease serine 2 (TMPRSS2), and basigin (BSG) as viral attack receptors and obtained PPI data from bioinformatics and system biology databases. They identified 59 proteins that interact with SARS-CoV-2 proteins and using the identified proteins they collected diseases named as comorbidities from DisGeNet [21], OMIM [22], ClinVar [23], and PheGenI [24] databases. They also selected 79 diseases from a list of diseases that are associated with 30 proteins selected from 59 proteins and the selections are not clear.
In [25], the authors explored the gene expression patterns from 30 array-based acute, chronic, and infectious gene expression datasets by meta-analysis. They observed the genes of the diseases are correlated and using literature mining they identified 78 genes which include receptors, and proteases of SARS-CoV-2 pathogenesis. Their findings also indicated that COVID-19 is susceptible to leukemia, nonalcoholic fatty liver disease, psoriasis, diabetes, and pulmonary arterial hypertension as comorbidities, through co-expression analysis of genes, pathways enrichment analysis, and significant enrichment of pathways.
Using data from preexisting diagnoses and hospitalized COVID-19 patients, the authors of [26] identified COVID-19 risk factors using the UK Biobank. Using logistic regression model, they estimated risks for each comorbidity from the existing comorbidities and found that males, people of black ethnicity, and no educational qualification people have a higher risk of COVID-19.
A study by the authors of [27] attempted to relate COVID-19 to breast cancer, colon cancer, kidney cancer, liver cancer, bladder cancer, prostate cancer, thyroid cancer, and lung cancer. Their findings identified dysregulated genes, pathways, gene ontologies, and associations and interactions between cancers and COVID-19. They marked the cancers as comorbidities based on their association and interaction with COVID-19. Anteriorly they selected the diseases, then COVID-19 is associated with the selected diseases.
In [28], the authors selected five illnesses: kidney, liver, diabetes, lung, and cardiovascular illness from the literature as comorbidities and examined their potential association with COVID-19. First, they identified the common genes of the comorbidities, then they conducted phenotype enrichment analysis and pathway enrichment analysis using the shared genes; they also identified common pathways between COVID-19 and the comorbidities and investigated the association of the pathways with COVID-19 itself.
By the authors of [29], laboratory-confirmed COVID-19 studies were meta-analyzed. In COVID-19 patients, the spread of comorbidities was evaluated, and the risk factor was identified. They found that older and male patients are more susceptible to COVID-19; also, they marked hypertension, diabetes, cardiovascular and followed by respiratory system diseases as prevalent risk factors from the literature.
Based on pooled proportion, the authors of [30] performed a systematic review and meta-analysis of comorbidities prevalence, severity, and mortality concerning age, gender, and geographic areas. They observed a high prevalence of comorbidities in the USA, the highest severity in Asia with the highest mortality in Europe and Latin American region; chronic kidney disease is highly responsible for disease severity and cerebrovascular accident, chronic kidney disease, and cardiovascular disease for mortality of COVID-19 patients. By reviewing the articles, they also got higher severity in female patients and mortality in older and male patients.
We analyzed the COVID-19 dataset in a systematic way to figure out the comorbidity.
P-value combination is very common in bioinformatics [31] and there are diverse methods for combining P-values from multiple statistical tests such as Brown’s Method [31], Fisher’s Method [32], Truncated Product Method [33], Stouffer’s Method [34], Tippett’s Method [35], Bonferroni Method [36], Weighted Z-test [37], Z-transform Test [38], Liptak Test [39], Sidak Test [40], Simes’ Test [40], Gamma Distribution [41] etc. For different situations and datasets, different methods are powerful and suitable [41]; in our case, Tippett’s Method is decent, and using Tippett’s Method we got COVID-19’s P-value. In later sections, a discussion about COVID-19’s P-value is there and after getting the P-value we did our further analysis.
3. Methods and Analysis
3.1. Covid-19 Dataset and Geo2r Analysis
For our investigation, we collected the dataset from National Center for Biotechnology Information (NCBI) (https://www.ncbi.nlm.nih.gov/) with accession number GSE166552 which is a microarray dataset prepared from blood samples of COVID-19 patients and healthy people. Initially, we analyzed the gene expression dataset with a web-based GEO2R tool, setting force normalization [42], precision weight (vooma) [43], and Benjamini & Hochberg (False discovery rate) [44] and comparing the transcriptomic profile of infected and control samples incorporating GEOquery [45], limma [46], and umap [47] R packages. The dataset comprises columns such as gene ID, adjusted P-value, P-value, Log2 fold change (logFC), gene sequence, and gene symbol. It was observed that certain cells within the gene symbol column were identified as missing values. Consequently, data related to these missing gene symbols were filtered out during the preprocessing phase. After preprocessing the dataset, we worked on the following analysis.
3.2. Dysregulated Genes and Disease Collection
To identify the comorbidities, statistically significant dysregulated (up-regulated and down-regulated) genes are selected as follows:
$$DGs = \left\{ \begin{array}{ll} \text{Up-regulated, } P\text{-value} \leq 0.05 \ \& \ \log FC \geq 1.0 \\ \text{Down-regulated, } P\text{-value} \leq 0.05 \ \& \ \log FC \leq 1.0 \end{array} \right.\tag{1}$$
Afterwards, more than 20,000 diseases associated with the selected DGs were obtained from several Gold Benchmark databases, including DisGeNET [21], ClinVar 2019 [23], PheWeb 2019 [48], dbGap [49], OMIM Disease [50], OMIM Expanded [50], Rare Diseases AutoRIF ARCHS4 Predictions [51], Rare Diseases GeneRIF ARCHS4 Predictions [51], Rare Diseases GeneRIF Gene Lists [51], Rare Diseases AutoRIF Gene Lists [51] and GWAS Catalog 2019 [52].
3.3. Covid-19’s P-value Calculation
The dataset used in this study was derived from COVID-19 patients and comprises a large number of genes with individual P-values. Our objective is to obtain a single P-value from the dataset representing the P-value of COVID-19. Thus, we combined all the genes’ P-values to get a single P-value that is considered COVID-19’s P-value using Tippett’s Method as follows:
$$p_c = 1 – \left(1 – \min(p_1, p_2, p_3, \ldots, p_i)\right)^k\tag{2}$$
Where min is the minimum function of P1, P2, P3….Pn as well as P1, P2, P3….Pn denote separate P-values and k is the total number of P-values of the genes of the dataset.
3.4. Euclidean Distance and Disease Order
In our investigation, we calculated the Euclidean distance between the P-value of COVID-19 and all the other individual diseases that were collected using the dysregulated genes. The distance is the absolute value difference of the P-values as follows:
$$E_d = |X_c – Y_i|$$
where Xc denotes the COVID-19’s P-value and Yi = Y1, Y2, Y3….Yi denotes the certain P-value of the collected disease. After getting the Euclidean distance of all other collected diseases from COVID-19, we sorted the collected diseases in ascending order depending on the Euclidean distance.
3.5. Clustering and Comorbidities Selection
In the collected diseases we found that some diseases have the same P-value and as a result, the same Euclidean distance is found for those diseases. We clustered the disease based on the same Euclidean distance of the sorted diseases list from top to bottom maintaining the ascending order. Finally, we select the comorbidities of COVID-19 from the top six clusters.
3.6. Our Proposed Model as An Algorithm
Algorithm: The algorithmic pseudocode to generate the workflows for identification of the comorbidities of COVID-19 as follows: | ||||
Input: Microarray dataset with six samples where three samples from COVID-19 patients and another three samples from healthy people. | ||||
Output: Related comorbidities of COVID-19. | ||||
· | Assign patient’s samples to the Case and healthy people’s samples to the Control group in the GEO2R tool. | |||
· | Analyze the dataset using GEO2R by setting force normalization, precision weights, and the Benjamini & Hochberg algorithm. | |||
· | Preprocess the dataset after analyzing it by GEO2R. | |||
· | Identify up-regulated genes applying . \(P\text{-}value \leq 0.05 \ \& \ \log FC \geq 1.0. | |||
· | Identify down-regulated genes applying . \(P\text{-}value \leq 0.05 \ \&\ \log FC \leq 1.0. | |||
· | Enter the identified dysregulated (up-regulated and down-regulated) genes into the Gold Benchmark databases and gather a list of diseases. | |||
· | Calculate COVID-19’s P-value using Tippett’s method. | |||
· | Calculate Euclidean distance from COVID-19 to all other diseases on the list. | |||
· | Sort all diseases of the list based on the Euclidean distance in ascending order. | |||
· | Cluster all the diseases on the list based on the same Euclidean distance. | |||
· | Select correlated comorbidities from the top clusters. | |||
end | ||||
The workflow steps of our proposed model are shown in Figure 1.
4. Results and Discussion
After analyzing, preprocessing, and filtering we got 35,011 differentially expressed genes with P-value, adjusted P-value, and fold change value. Then we evicted 1,421 up-regulated genes using a statistical threshold of \(P\text{-}value \leq 0.05\) and absolute Log2 fold change \(log FC \geq 1.0\) . Also, we got 2,138 down-regulated genes using \(P\text{-}value \leq 0.05\) and Log2 fold change \(log FC \leq 1.0\) ; a total of 3,559 dysregulated genes were identified. To attain the result, we uploaded a total of 3,559 dysregulated genes in the stated method and analysis section’s eleven Gold Benchmark databases of Enrichr [53] which is a web-based gene set enrichment analyzing tool, and got a total of 22,820 common and rare type diseases with P-value, adjusted P-value, gene symbol and many other attributes. Simultaneously we got the P-value of COVID-19 as 0.301041817 by combining the P-values of all 35,011 genes from our filtered dataset using Tippett’s method. Next Euclidean distances among the P-value of COVID-19 and 22,820 diseases are enumerated individually and then the total of 22,820 diseases are rearranged in ascending order using the computed Euclidean distances.
Table 1 shows the primary disease and its P-value, top collected diseases, P-values of the top collected diseases, and Euclidean distance from the primary disease to the top collected diseases from left to right. After sorting the collected disease, we got prominent ear, acute myelocytic leukemia, adducted thumb, cancer of urinary tract, increased appetite, malignant neoplasm of vulva, myelodysplastic syndrome with isolated del(5q), nonnuclear polymorphic congenital cataract, chromosome 12 12p trisomy, chromosome 15q trisomy, chromosome 9 monosomy 9p, body weight changes, abdominal aortic aneurysm, kidney neoplasm, diabetes mellitus, and other diseases from top of the total diseases. Many diseases have the same Euclidean distance from COVID-19 and diseases with similar Euclidean distances are retained in the same cluster.
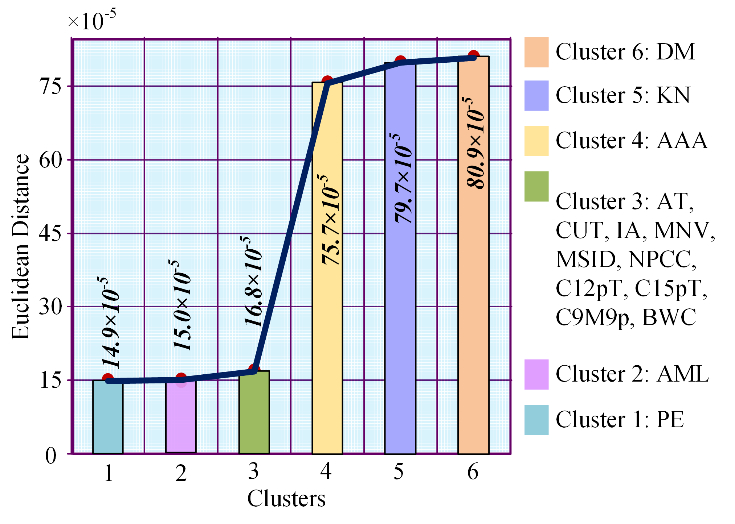
Table 2 represents cluster number, Euclidean distance between COVID-19 and the clusters or clusters’ disease, and the disease names in the clusters from left to right. Euclidean distance between cluster 1 and COVID-19 is 0.000149032; in the same way, Euclidean distances are 0.0001505, 0.000168149, 0.000756942, 0.000798617 and 0.000808946 among COVID-19 and second cluster, third cluster, fourth cluster, fifth cluster, and sixth cluster.
We clustered the entire 22,820 diseases based on the same Euclidean distance and took the top six clusters. From Figure 2, we see that in cluster no. 1 there is only one disease prominent ear; in cluster 2, only acute myelocytic leukemia is present; cluster 3 contains ten diseases and they are adducted thumb, cancer of urinary tract, increased appetite, malignant neoplasm of vulva, myelodysplastic syndrome with isolated del(5q), nonnuclear polymorphic congenital cataract, chromosome 12 12p trisomy, chromosome 15q trisomy, chromosome 9 monosomy 9p and body weight changes. Again, cluster 4, cluster 5, and cluster 6 hold abdominal aortic aneurysm, kidney neoplasm, and diabetes mellitus.
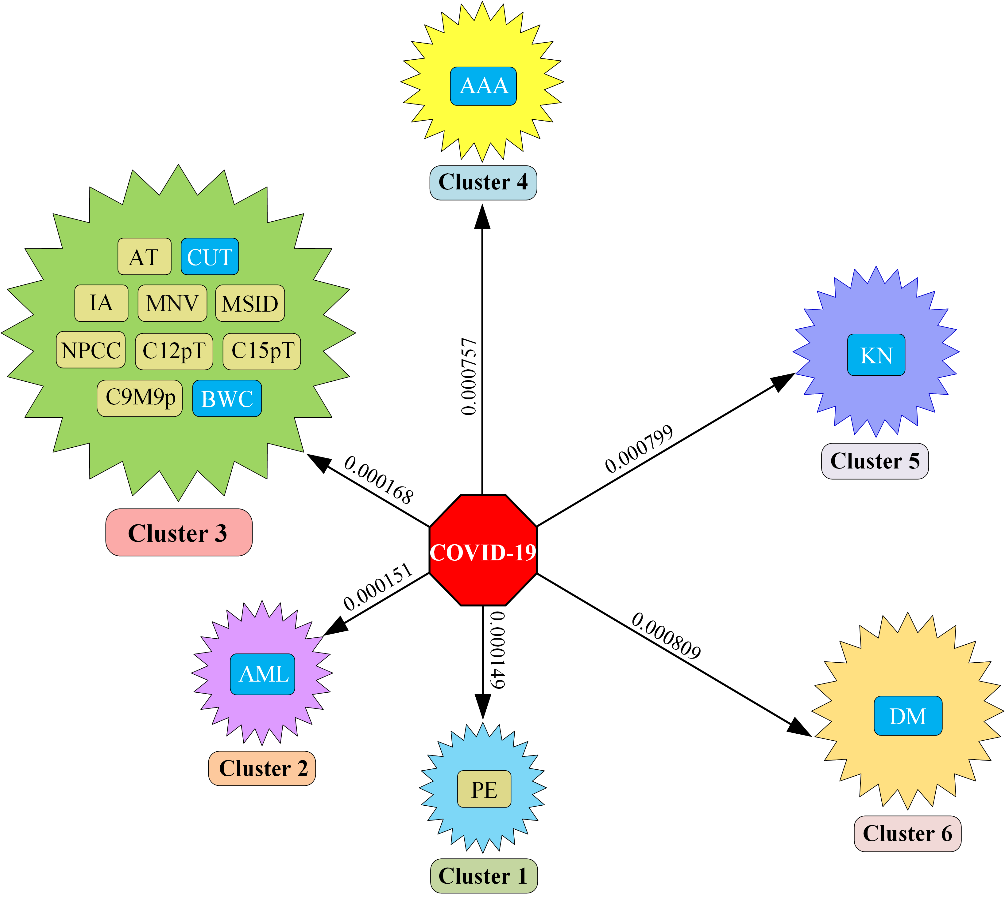
Finally, we selected comorbidity from our top six clusters. Figure 3 shows that acute myelocytic leukemia is selected as comorbidity from cluster 2, from cluster 3 cancer of urinary tract and body weight changes are selected as comorbidity; abdominal aortic aneurysm, kidney neoplasm, and diabetes mellitus are selected from cluster 4, cluster 5, and cluster 6; only cluster 1 is not considered to select comorbidity as we have not found a relationship between COVID-19 and prominent ear in global research.
We developed a computational and bioinformatics framework to identify the comorbidities of COVID-19 and our model is suitable to obtain the comorbidities of other diseases quickly. As far as we know there is no other such approach to ascertain comorbidities of diseases numerically; there are clinical tests, meta-analysis, and cross-checking tests to determine comorbidities but they are time-consuming and take much exertion. Acute myelocytic leukemia [54] — [57], cancer of urinary tract [58], [59], body weight changes [60] [61], abdominal aortic aneurysm [62] [63], kidney neoplasm [64] — [66], and diabetes mellitus [67] — [70] are the outcomes of our research where cancer of urinary tract and kidney neoplasm are related to bladder cancer [71], [72]. Moreover, bladder cancer is a post COVID-19 condition [59]. So, there is a chance of ensuing cancer of urinary tract and kidney neoplasm in COVID-19 recovered patients. Body weight changes (weight gain or weight loss) is also a post COVID-19 condition [60], [73] and excess weight gain or overweight causes obesity [74].
What we found in our outcomes is also the same in the published literature; thus, we validated our work.
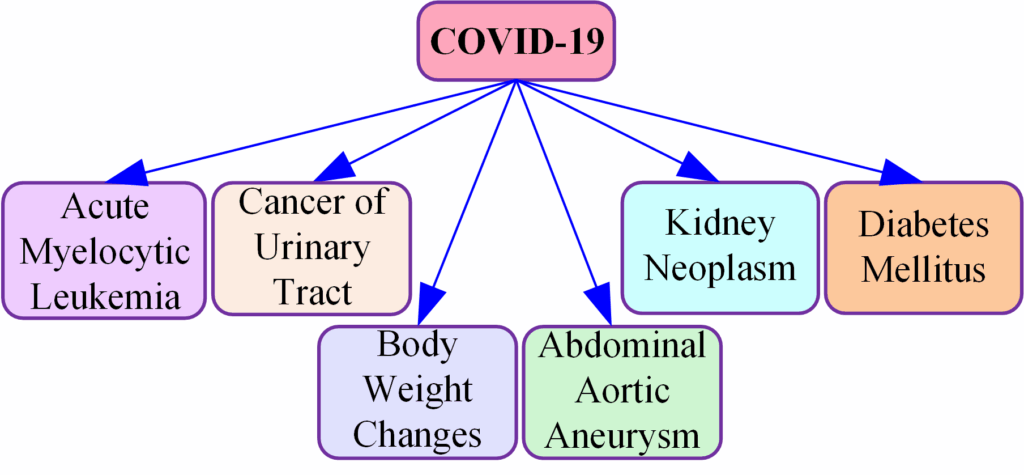
Clinical decision-making and outcomes are significantly affected by comorbidities, which influence treatment strategies and patient management. Research indicates that comorbid conditions complicate the treatment landscape by influencing drug interactions, therapeutic efficacy, and patient compliance [75]. Additionally, comorbidity can complicate the prognosis of primary illnesses, increasing healthcare expenditures [76]. Early detection of comorbidities enables healthcare providers to implement preventive measures and targeted interventions to arrest disease progression and improve patient health [77]. It is also essential to assess and optimize patients with comorbidities before surgery to reduce complications during surgery [78]. Therefore, it is imperative to identify accurate comorbidities early in clinical practice to optimize treatment strategies, improve patient management, and improve health outcomes. Comorbidities that are the result of our work are shown in Figure 4.
Table 1: Summary of results along with the P-value of COVID-19, top collected disease, their P-values
and Euclidean distance between COVID-19 and the top collected disease
Primary Disease | P-value of COVID-19 | Top Collected Disease | Collected Disease’s P-value | Euclidean Distance |
COVID-19 | 0.301041817 | Prominent Ear | 0.300892784 | 0.000149032 |
Acute Myelocytic Leukemia | 0.301192317 | 0.0001505 | ||
Adducted Thumb | 0.301209965 | 0.000168149 | ||
Cancer of Urinary Tract | 0.301209965 | 0.000168149 | ||
Increased Appetite | 0.301209965 | 0.000168149 | ||
Malignant Neoplasm of Vulva | 0.301209965 | 0.000168149 | ||
Myelodysplastic Syndrome with Isolated del(5q) | 0.301209965 | 0.000168149 | ||
Nonnuclear Polymorphic Congenital Cataract | 0.301209965 | 0.000168149 | ||
Chromosome 12 12p Trisomy | 0.301209965 | 0.000168149 | ||
Chromosome 15q Trisomy | 0.301209965 | 0.000168149 | ||
Chromosome 9 Monosomy 9p | 0.301209965 | 0.000168149 | ||
Body Weight Changes | 0.301209965 | 0.000168149 | ||
Abdominal Aortic Aneurysm | 0.300284875 | 0.000756942 | ||
Kidney Neoplasm | 0.3002432 | 0.000798617 | ||
Diabetes Mellitus | 0.301850763 | 0.000808946 |
Table 2: Disease in each cluster and Euclidean distance from COVID-19 to clusters or clusters’ disease
No. | Euclidean distance | Diseases in the cluster |
Cluster 1 | 0.000149032 | Prominent Ear |
Cluster 2 | 0.0001505 | Acute Myelocytic Leukemia |
Cluster 3 | 0.000168149 | Adducted Thumb |
Cancer of Urinary Tract | ||
Increased Appetite | ||
Malignant Neoplasm of Vulva | ||
Myelodysplastic Syndrome with Isolated del(5q) | ||
Nonnuclear Polymorphic Congenital Cataract | ||
Chromosome 12 12p Trisomy | ||
Chromosome 15q Trisomy | ||
Chromosome 9 Monosomy 9p | ||
Body Weight Changes | ||
Cluster 4 | 0.000756942 | Abdominal Aortic Aneurysm |
Cluster 5 | 0.000798617 | Kidney Neoplasm |
Cluster 6 | 0.000808946 | Diabetes Mellitus |
5. Conclusion
COVID-19’s comorbidities increase the disease severity and mortality as well as influence organ damage of the patients; often the patients die due to their main comorbidities. Because of that, it is exigent for medical practitioners to find out the comorbidities of diseases, and in our study, we developed a pipeline that can detect comorbidities of COVID-19 from genetic datasets in a computational and bioinformatics way. We identified and diseases related to the using the bioinformatics approach as well as detected risk factor diseases linked with COVID-19 as comorbidities from the gathered diseases in a systematic computational way.
In this research, one of the main challenges for us is to find the accurate P-value of COVID-19. We observed that the more accurate COVID-19’s P-value gives the more accurate comorbidities as a result. In the future, we will apply other alternative methods to derive the exact P-value of COVID-19. We suggest that our developed approach will assist in the diagnosis of comorbidities of other diseases early if the genetic dataset is available and thus this model will help healthcare systems to reduce the comorbidity diagnosis cost of a disease.
Conflict of Interest
The authors declare no conflict of interest.
Acknowledgment
The assistance for this project is provided by the ICT Division of the Government of the People’s Republic of Bangladesh, under Grant number 22FS15614.
- T. Acter, N. Uddin, J. Das, A. Akhter, “Since January 2020 Elsevier has created a Covid-19 resource centre with free information in English and Mandarin on the novel coronavirus Covid- 19 . The Covid-19 resource centre is hosted on Elsevier Connect , the company ’ s public news and information,” Science of the Total Environment Journal, vol. N/V, no. January, pp. 2–15, 2020.
- Coronavirus disease (COVID-19) (no date) Who.int. Available at: https://www.who.int/health-topics/coronavirus (Accessed: June 20, 2021).
- W.J. Guan, W.H. Liang, J.X. He, N.S. Zhong, “Cardiovascular comorbidity and its impact on patients with COVID-19,” European Respiratory Journal, vol. 55, no. 6, pp. 1069–1076, 2020, doi:10.1183/13993003.01227-2020.
- M.J. Hasan, A.M. Anam, S.M.R. Huq, R. Rabbani, “Impact of Comorbidities on Clinical Outcome of Patients with COVID-19: Evidence from a Single-center in Bangladesh,” Health Scope, vol. 10, no. 1, pp. 1–9, 2021, doi:10.5812/jhealthscope.109268.
- H. Ejaz, A. Alsrhani, A. Zafar, H. Javed, K. Junaid, A.E. Abdalla, K.O.A. Abosalif, Z. Ahmed, S. Younas, “COVID-19 and comorbidities: Deleterious impact on infected patients,” Journal of Infection and Public Health, vol. 13, no. 12, pp. 1833–1839, 2020, doi:10.1016/j.jiph.2020.07.014.
- Y. Bai, L. Yao, T. Wei, F. Tian, D.Y. Jin, L. Chen, M. Wang, “Presumed asymptomatic carrier transmission of COVID-19” Jama, vol. 323, no. 14, pp.1406-1407, 2020.
- C. Huang, Y. Wang, X. Li, L. Ren, J. Zhao, Y. Hu, L. Zhang, G. Fan, J. Xu, X. Gu, Z. Cheng, T. Yu, J. Xia, Y. Wei, W. Wu, X. Xie, W. Yin, H. Li, M. Liu, Y. Xiao, H. Gao, L. Guo, J. Xie, G. Wang, R. Jiang, Z. Gao, Q. Jin, J. Wang, B. Cao, “Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China,” The Lancet, vol. 395, no. 10223, pp. 497–506, 2020, doi:10.1016/S0140-6736(20)30183-5.
- N. Chen, M. Zhou, X. Dong, J. Qu, F. Gong, Y. Han, Y. Qiu, J. Wang, Y. Liu, Y. Wei, J. Xia, T. Yu, X. Zhang, L. Zhang, “Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study,” The Lancet, vol. 395, no. 10223, pp. 507–513, 2020, doi:10.1016/S0140-6736(20)30211-7.
- Q. Gao, Y. Hu, Z. Dai, F. Xiao, J. Wang, J. Wu, “The Epidemiological Characteristics of 2019 Novel Coronavirus Diseases (COVID-19) in Jingmen, China,” SSRN Electronic Journal, vol. 2, no. 8, pp. 113–122, 2020, doi:10.2139/ssrn.3548755.
- M. Fathi, K. Vakili, F. Sayehmiri, A. Mohamadkhani, M. Hajiesmaeili, M. Rezaei-Tavirani, O. Eilami, “The prognostic value of comorbidity for the severity of COVID-19: A systematic review and meta-analysis study,” PloS one, vol. 16, no. 2, p.e0246190, 2021.
- X. Yu, Y. Zhao, Y. Yang, W. He, N. Yang, F. Su, J. Zhu, Z. Zhu, “The magnetic and electronic properties of Ho6Fe23−xCox (x = 0, 1, 2, 3),” Journal of Magnetism and Magnetic Materials, vol. 597, , 2024, doi:10.1016/j.jmmm.2024.172029.
- B. Wang, R. Li, Z. Lu, Y. Huang, “Does comorbidity increase the risk of patients with covid-19: Evidence from meta-analysis,” Aging, vol. 12, no. 7, pp. 6049–6057, 2020, doi:10.18632/AGING.103000.
- P. Qiu, Y. Zhou, F. Wang, H. Wang, M. Zhang, X. Pan, Q. Zhao, J. Liu, “Clinical characteristics, laboratory outcome characteristics, comorbidities, and complications of related COVID-19 deceased: a systematic review and meta-analysis,” Aging Clinical and Experimental Research, vol. 32, no. 9, pp. 1869–1878, 2020, doi:10.1007/s40520-020-01664-3.
- A. Baradaran, M.H. Ebrahimzadeh, A. Baradaran, A.R. Kachooei, “Prevalence of comorbidities in COVID-19 patients: A systematic review and meta-analysis,” Archives of Bone and Joint Surgery, vol. 8, no. SpecialIssue, pp. 247–255, 2020, doi:10.22038/abjs.2020.47754.2346.
- A. Zaenab, V. Jose, C. J. Cynthia, Z. Yousuf, M. Claudia, S. Hamed, “The Effects of Co-morbidities on COVID-19 Patients Admitted to the Hospital,” Fam Med Med Sci Res 10:2, 2021, doi: 10.35248/2327-4972.21.10.261.
- WHO Coronavirus (COVID-19) Dashboard (no date) Who.int. Available at: https://covid19.who.int/ (Accessed: June 22, 2021).
- A. Clark, M. Jit, C. Warren-Gash, B. Guthrie, H.H.X. Wang, S.W. Mercer, C. Sanderson, M. McKee, C. Troeger, K.L. Ong, F. Checchi, P. Perel, S. Joseph, H.P. Gibbs, A. Banerjee, R.M. Eggo, E.S. Nightingale, K. O’Reilly, T. Jombart, W.J. Edmunds, A. Rosello, F.Y. Sun, K.E. Atkins, N.I. Bosse, S. Clifford, T.W. Russell, A.K. Deol, Y. Liu, S.R. Procter, et al., “Global, regional, and national estimates of the population at increased risk of severe COVID-19 due to underlying health conditions in 2020: a modelling study,” The Lancet Global Health, vol. 8, no. 8, pp. e1003–e1017, 2020, doi:10.1016/S2214-109X(20)30264-3.
- M.F. Beckman, F.B. Mougeot, J.L.C. Mougeot, “Comorbidities and susceptibility to covid-19: A generalized gene set data mining approach,” Journal of Clinical Medicine, vol. 10, no. 8, 2021, doi:10.3390/jcm10081666.
- S. Aktar, A. Talukder, M.M. Ahamad, A.H.M. Kamal, J.R. Khan, M. Protikuzzaman, N. Hossain, A.K.M. Azad, J.M.W. Quinn, M.A. Summers, T. Liaw, V. Eapen, M.A. Moni, “Machine learning approaches to identify patient comorbidities and symptoms that increased risk of mortality in covid-19,” Diagnostics, vol. 11, no. 8, pp. 1–18, 2021, doi:10.3390/diagnostics11081383.
- B. Chakrabarty, D. Das, G. Bulusu, A. Roy, “Network-Based Analysis of Fatal Comorbidities of COVID-19 and Potential Therapeutics,” IEEE/ACM Transactions on Computational Biology and Bioinformatics, vol. 18, no. 4, pp. 1271–1280, 2021, doi:10.1109/TCBB.2021.3075299.
- J. Piñero, Á. Bravo, N. Queralt-Rosinach, A. Gutiérrez-Sacristán, J. Deu-Pons, E. Centeno, J. García-García, F. Sanz, L.I. Furlong, “DisGeNET: A comprehensive platform integrating information on human disease-associated genes and variants,” Nucleic Acids Research, vol. 45, no. D1, pp. D833–D839, 2017, doi:10.1093/nar/gkw943.
- J.S. Amberger, C.A. Bocchini, F. Schiettecatte, A.F. Scott, A. Hamosh, “OMIM.org: Online Mendelian Inheritance in Man (OMIM®), an Online catalog of human genes and genetic disorders,” Nucleic Acids Research, vol. 43, no. D1, pp. D789–D798, 2015, doi:10.1093/nar/gku1205.
- M.J. Landrum, J.M. Lee, M. Benson, G. Brown, C. Chao, S. Chitipiralla, B. Gu, J. Hart, D. Hoffman, J. Hoover, W. Jang, K. Katz, M. Ovetsky, G. Riley, A. Sethi, R. Tully, R. Villamarin-Salomon, W. Rubinstein, D.R. Maglott, “ClinVar: Public archive of interpretations of clinically relevant variants,” Nucleic Acids Research, vol. 44, no. D1, pp. D862–D868, 2016, doi:10.1093/nar/gkv1222.
- E.M. Ramos, D. Hoffman, H.A. Junkins, D. Maglott, L. Phan, S.T. Sherry, M. Feolo, L.A. Hindorff, “Phenotype-genotype integrator (PheGenI): Synthesizing genome-wide association study (GWAS) data with existing genomic resources,” European Journal of Human Genetics, vol. 22, no. 1, pp. 144–147, 2014, doi:10.1038/ejhg.2013.96.
- M. K. Singh, A. Mobeen, A. Chandra, S. Joshi, S. Rama-chandran, “A meta-analysis of comorbidities in COVID-19: Which diseases increase the susceptibility of SARS-CoV-2 infection?,” Computers in biology and medicine, 130, p.104219, 2021.
- J.L. Atkins, J.A.H. Masoli, J. Delgado, L.C. Pilling, C.L. Kuo, G.A. Kuchel, D. Melzer, “Preexisting Comorbidities Predicting COVID-19 and Mortality in the UK Biobank Community Cohort,” Journals of Gerontology – Series A Biological Sciences and Medical Sciences, vol. 75, no. 11, pp. 2224–2230, 2020, doi:10.1093/gerona/glaa183.
- S. Satu, I. Khan, R. Rahman, K.C. Howlader, S. Roy, S.S. Roy, J.M.W. Quinn, M.A. Moni, “Diseasome and comorbidities complexities of SARS-CoV-2 infection with common malignant diseases,” Briefings in Bioinformatics, vol. 22, no. 2, pp. 1415–1429, 2021, doi:10.1093/bib/bbab003.
- M.E. Dolan, D.P. Hill, G. Mukherjee, M.S. McAndrews, E.J. Chesler, J.A. Blake, “Investigation of COVID-19 comorbidities reveals genes and pathways coincident with the SARS-CoV-2 viral disease,” Scientific Reports, vol. 10, no. 1, pp. 1–11, 2020, doi:10.1038/s41598-020-77632-8.
- J. Yang, Y. Zheng, X. Gou, K. Pu, Z. Chen, Q. Guo, R. Ji, H. Wang, Y. Wang, Y. Zhou, “Prevalence of comorbidities and its effects in coronavirus disease 2019 patients: A systematic review and meta-analysis,” International Journal of Infectious Diseases, vol. 94, , pp. 91–95, 2020, doi:10.1016/j.ijid.2020.03.017.
- B. Thakur, P. Dubey, J. Benitez, J.P. Torres, S. Reddy, “OPEN A systematic review and meta ‑ analysis of geographic differences in comorbidities and associated severity and mortality among individuals with COVID ‑ 19,” Scientific Reports, pp. 1–13, 2021, doi:10.1038/s41598-021-88130-w.
- W. Poole, D.L. Gibbs, I. Shmulevich, B. Bernard, T.A. Knijnenburg, “Combining dependent P- values with an empirical adaptation of Brown ’ s method,” no. 2002, pp. 430–436, 2016, doi:10.1093/bioinformatics/btw438.
- Q. Li, J. Ding, “Fisher ’ s method of combining dependent statistics using generalizations of the gamma distribution with applications to genetic pleiotropic associations,” pp. 284–295, 2014, doi:10.1093/biostatistics/kxt045.
- D. V Zaykin, L.A. Zhivotovsky, P.H. Westfall, B.S. Weir, “Truncated Product Method for Combining P-Values,” vol. 185, no. November 2000, pp. 170–185, 2002, doi:10.1002/gepi.0042.
- N.A. Heard, P. Rubin-delanchy, “Choosing Between Methods of Combining p -values arXiv : 1707 . 06897v4 [ stat . ME ] 14 Dec 2017,” pp. 1–13, 2017.
- L.H.C. Tippett, “The methods of statistics,” The Meth-ods of Statistics, pp.222, ref. 66, 1931.
- V. Vovk, S.T. Oct, “Combining p-values via averaging,” 2019.
- D. V Zaykin, “Optimally weighted Z -test is a powerful method for combining probabilities in meta-analysis,” vol. 24, , pp. 1836–1841, 2011, doi:10.1111/j.1420-9101.2011.02297.x.
- M.C. Whitlock, “Combining probability from independent tests : the weighted Z -method is superior to Fisher ’ s approach,” vol. 18, , pp. 1368–1373, 2005, doi:10.1111/j.1420-9101.2005.00917.x.
- F. Solmi, P. Onghena, “Combining p-values in replicated single-case experiments with multivariate outcome,” vol. 24, , pp. 607–633, 2014.
- V.C. Tests, “Combining P Values to Get More Power,” 2005, doi:10.1002/0470011815.b2a15181.
- Z. Chen, W. Yang, Q. Liu, J.Y. Yang, J. Li, M.Q. Yang, “A new statistical approach to combining p-values using gamma distribution and its application to genome-wide association study,” vol. 15, no. Suppl 17, pp. 1–7, 2014.
- T.P. Speed, B.M. Bolstad, R.A. Irizarry, M. Astrand, “A comparison of normalization methods for high density oligonucleotide array data based on variance and bias,” vol. 19, no. 2, pp. 185–193, 2003.
- C.W.M. Law, “Precision weights for gene expression analysis (Doctoral dissertation),” 2013.
- N.K. Podder, S. Babu, S. Omit, P.C. Shill, H.K. Rana, “Genetic Effects of Covid 19 on the Development of Neurodegenerative Diseases,” no. December 2021, pp. 17–19, 2022.
- S. Davis, P.S. Meltzer, “BIOINFORMATICS APPLICATIONS NOTE GEOquery : a bridge between the Gene Expression Omnibus ( GEO ) and BioConductor,” vol. 23, no. 14, pp. 1846–1847, 2007, doi:10.1093/bioinformatics/btm254.
- G.K. Smyth, “Limma : Linear Models for Microarray Data,” no. 2005, pp. 397–420.
- T. Uniform, M. Approximation, I. Matrix, L. Rcpp, “Package ‘ umap ,’” pp. 1–7, 2023.
- F. Shahabinezhad, P. Mosaddeghi, M. Negahdaripour, M. Farahmandnejad, M.J. Taghipour, M. Moghadami, N. Nezafat, S.M. Masoompour, “Therapeutic approaches for COVID-19 based on the dynamics of interferon- mediated immune responses,” no. March, 2020, doi:10.20944/preprints202003.0206.v2.
- H. Rahman, H.K. Rana, S. Peng, X. Hu, C. Chen, J.M.W. Quinn, M.A. Moni, “Bioinformatics and machine learning methodologies to identify the effects of central nervous system disorders on glioblastoma progression Bioinformatics and machine learning methodologies to identify the effects of central nervous system disorders on glio,” no. January, 2021, doi:10.1093/bib/bbaa365.
- Z. Nain, H.K. Rana, P. Liò, S. Mohammed, S. Islam, M.A. Summers, “Pathogenetic profiling of COVID-19 and SARS-like viruses,” vol. 22, no. 2, pp. 1175–1196, 2021, doi:10.1093/bib/bbaa173.
- G. V Glinsky, “Impacts of genomic networks governed by human-specific regulatory sequences and genetic loci harboring fixed human-specific neuro-regulatory single nucleotide mutations on phenotypic traits of modern humans,” Chromosome Research, vol.28, pp. 331–354, 2020, doi: 10.1007/s10577-020-09639-w
- P. Mosaddeghi, F. Shahabinezhad, Z. Dehghani, M. Farahmandnejad, M. J. Taghipour, M. Moghadami, N. Nezafat, S. M. Masoompour, M. Negahdaripour, “Therapeutic Approaches for COVID-19 Based on the Interferon-mediated Immune Responses”, Current Signal Transduction Therapy, vol. 16, no. 1. 2021, doi:10.2174/1574362416666210120104636
- M. V Kuleshov, M.R. Jones, A.D. Rouillard, N.F. Fernandez, Q. Duan, Z. Wang, S. Koplev, S.L. Jenkins, K.M. Jagodnik, A. Lachmann, M.G. Mcdermott, C.D. Monteiro, W. Gundersen, A. Ma, “Enrichr : a comprehensive gene set enrichment analysis web server 2016 update,” vol. 44, no. May, pp. 90–97, 2016, doi:10.1093/nar/gkw377.
- A.M. Khan, Z. Ajmal, M. Raval, E. Tobin,“Con-current diagnosis of acute myeloid leukemia and COVID-19: a man-agement challenge,” Cureus, vol. 12, no. 8, 2020, doi:10.7759/cureus.9629
- H.A. Tehrani, M. Darnahal, S.A. Nadji, S. Haghighi, “COVID-19 re-infection or persistent infection in patient with acute myeloid leukaemia M3 : a mini review,” New Microbes and New Infections, vol. 39, , pp. 100830, 2021, doi:10.1016/j.nmni.2020.100830.
- A.O. Brien, J. Campling, H. Goodman, C.L. Chang, “Respirology Case Reports presentation and management implications,” vol. 8, , pp. 1–5, 2020, doi:10.1002/rcr2.650.
- M. Gavillet, J.C. Klappert, O. Spertini, S. Blum, “Acute leukemia in the time of COVID-19,” Leukemia research, 92, p.106353, 2020.
- J. Sarkis, R. Samaha, “Bladder cancer during the COVID-19 pandemic : the calm before the storm ?,” Future Science, vol. 6, no. 8, pp. 13–15, 2020, doi: 10.2144/fsoa-2020-0101
- The experience of UK patients with bladder cancer during the COVID-19 pandemic : a survey-based snapshot, pp. 179–181, 2021, doi:10.1111/bju.15287.
- L. Di Filippo, R. De Lorenzo, E. Cinel, E. Falbo, M. Ferrante, M. Cilla, S. Martinenghi, G. Vitali, E. Bosi, A. Giustina, P. Rovere-Querini, C. Conte, “Weight trajectories and abdominal adiposity in COVID-19 survivors with overweight/obesity,” International Journal of Obesity, vol. 45, no. 9, pp. 1986–1994, 2021, doi:10.1038/s41366-021-00861-y.
- H.S.J. Chew, V. Lopez, “Global impact of covid-19 on weight and weight-related behaviors in the adult population: A scoping review,” International Journal of Environmental Research and Public Health, vol. 18, no. 4, pp. 1–32, 2021, doi:10.3390/ijerph18041876.
- M. Shih, B. Swearingen, R. Rhee, N. York, “Since January 2020 Elsevier has created a COVID-19 resource centre with free information in English and Mandarin on the novel coronavirus COVID- 19 . The COVID-19 resource centre is hosted on Elsevier Connect , the company ’ s public news and information ,” no. January, 2020.
- M.H. Syed, M. Wheatcroft, D. Marcuzzi, H. Hennessey, M. Qadura, “Management of a mycotic aneurysm in a patient with COVID-19: A case report,” Medicina (Lithuania), vol. 57, no. 6, pp. 1–7, 2021, doi:10.3390/medicina57060620.
- I. Tsimafeyeu, G. Alekseeva, M. Berkut, A. Nosov, I. Myslevtsev, A. Andrianov, A. Semenov, P. Borisov, R. Zukov, V. Goutnik, S. Savchuk, M. Volkova, M. Mukhina, “COVID-19 in Patients With Renal Cell Carcinoma in the Russian Federation,” Clinical Genitourinary Cancer, vol. 19, no. 2, pp. e69–e71, 2021, doi:10.1016/j.clgc.2020.07.007.
- M. Mihalopoulos, N. Dogra, N. Mohamed, K. Badani, N. Kyprianou, “COVID-19 and Kidney Disease: Molecular Determinants and Clinical Implications in Renal Cancer,” European Urology Focus, vol. 6, no. 5, pp. 1086–1096, 2020, doi:10.1016/j.euf.2020.06.002.
- N. Zaki, H. Alashwal, S. Ibrahim, “Association of hypertension, diabetes, stroke, cancer, kidney disease, and high-cholesterol with COVID-19 disease severity and fatality: A systematic review,” Diabetes and Metabolic Syndrome: Clinical Research and Reviews, vol. 14, no. 5, pp. 1133–1142, 2020, doi:10.1016/j.dsx.2020.07.005.
- C. Sardu, G. Gargiulo, G. Esposito, G. Paolisso, R. Marfella, “Impact of diabetes mellitus on clinical outcomes in patients affected by Covid-19,” Cardiovascular Diabetology, vol. 19, no. 1, pp. 4–7, 2020, doi:10.1186/s12933-020-01047-y.
- L. Fang, G. Karakiulakis, M. Roth, “Are patients with hypertension and diabetes mellitus at increased risk for COVID-19 infection?,” The Lancet Respiratory Medicine, vol. 8, no. 4, pp. e21, 2020, doi:10.1016/S2213-2600(20)30116-8.
- S. Lim, J.H. Bae, H.S. Kwon, M.A. Nauck, “COVID-19 and diabetes mellitus: from pathophysiology to clinical management,” Nature Reviews Endocrinology, vol. 17, no. 1, pp. 11–30, 2021, doi:10.1038/s41574-020-00435-4.
- S.B. Sen Omit, S. Akhter, H.K. Rana, A.R.M.M.H. Rana, N.K. Podder, M.I. Rakib, A. Nobi, “Identification of Comorbidities, Genomic Associations, and Molecular Mechanisms for COVID-19 Using Bioinformatics Approaches,” BioMed Research International, vol. 2023, no. December 2019, 2023, doi:10.1155/2023/6996307.
- A.F. Kantor, P. Hartge, R.N. Hoover, A.S. Narayana, J.W. Sullivan, J.F. Fraumeni, “Urinary tract infection and risk of bladder cancer,” American Journal of Epidemiology, vol. 119, no. 4, pp. 510–515, 1984, doi:10.1093/oxfordjournals.aje.a113768.
- T. Brown, R. Slack, L. Rushton, “Occupational cancer in Britain: Urinary tract cancers: Bladder and kidney,” British Journal of Cancer, vol. 107, no. S1, pp. S76–S84, 2012, doi:10.1038/bjc.2012.121.
- C.D.S. Costa, E.M. Steele, M.A. Leite, F. Rauber, R.B. Levy, C.A. Monteiro, “Body weight changes in the NutriNet Brasil cohort during the covid-19 pandemic,” Revista de Saude Publica, vol. 55, , pp. 1–5, 2021, doi:10.11606/S1518-8787.2021055003457.
- D.C. Van Duijvenbode, M.J.M. Hoozemans, M.N.M. Van Poppel, K.I. Proper, “The relationship between overweight and obesity, and sick leave: A systematic review,” International Journal of Obesity, vol. 33, no. 8, pp. 807–816, 2009, doi:10.1038/ijo.2009.121.
- L. Smith, J. Taylor, A. Lewis, “Tailored Medication Regimens for Patients with Comorbidities: A Review,” Journal of Pharmacy Practice, 32(5), 561–569, 2019.
- A. L. Jones, P. S. Duggan, S. O’Brien, “Comorbidities and Mental Health Disorders: The Need for Integrated Care,” Irish Medical Journal, 113(6), 89, 2020.
- A. Marengoni, S. Angleman, R. Melis, F. Mangialasche, A. Karp, A. Garmen, B. Meinow, L. Fratiglioni, “Aging with multimorbidity: A systematic review of the literature,” Ageing Research Reviews, vol. 10, no. 4, pp. 430–439, 2011, doi:10.1016/j.arr.2011.03.003.
- E. A. Brown, E. Mahanna, F. J. Schiffman, “Comorbidity Assessment in Surgical Patients,” Anesthesiology Clinics, 36(1), 1–14, 2018.
- Shudeb Babu Sen Omit, Md Mohiuddin, Salma Akhter, Md. Hasan Imam, A. K. M. Mostofa Kamal Habib, Syed Mohammad Meraz Hossain, Nitun Kumar Podder, “COVID-19 Pandemic and Mental Well-Being: A Study Conducted on Medical Students and Their Parents in a Private Medical College in Pakistan”, Journal of Engineering Research and Sciences, vol. 2, no. 2, pp. 1–7, 2023. doi: 10.55708/js0202001
- Shudeb Babu Sen Omit, Md Mohiuddin, Salma Akhter, Md. Hasan Imam, A. K. M. Mostofa Kamal Habib, Syed Mohammad Meraz Hossain, Nitun Kumar Podder, “The Future of Work after COVID-19 from the Office to the Edge: Using the IT Industry as an Example”, Journal of Engineering Research and Sciences, vol. 1, no. 9, pp. 15–32, 2022. doi: 10.55708/js0109003
